Cladistic analysis allows biologists to trace evolutionary relationships among organisms by analyzing their characteristics. Sequence analysis made it possible to apply bioinformatics techniques to cladistics and has led to extraordinary discoveries about the history of life on earth.
Cladistics is the field of study in which biologists trace the evolutionary lineages of species. Each time a speciation even occurs and a
species splits into two species, each new species begins to acquire changes, both adaptive and incidental, that differentiate it from its sister species.
These speciations can be mapped out into a tree diagram called a cladogram. A clade is a branch on the tree diagram, and it includes an ancestor species and all its descendants.
Diagram Representation
Although cladograms are represented by tree diagrams, they could also be thought of as Euler diagrams in which all the sets are

completely nested within other sets. In these diagrams, each set has either zero or two subsets, depending on whether it spawned daughter species or not. Each subset, in turn, may each have zero or two subsets.
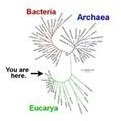
All cellular life on earth shares a common ancestor. The cladogram of all species descended from this ancestor, called the Last Universal Common Ancestor (LUCA), is known as the Tree of Life.
Above right: A cladogram of the tree of life, showing where our species falls. Source: NSF.gov
Directly right: Biological classification rendered as a modified Euler diagram, with each subrank nested within a higher rank. Image by Peter Halasz, licensed under CC Attribution ShareAlike 2.5 and CC Attribution ShareAlike Unported 3.0.
How Cladistic Analysis Works
Cladistics begins by selecting a group of species, then analyzing characteristics shared by members of the group. In cladistics, the emphasis is on keeping all groups monophyletic and avoiding groups that are paraphyletic or polyphyletic.
A monophyletic group includes all the descendants of a common ancestor and no species that did not descend from that ancestor.
A paraphyletic group contains only some descendants of a common ancestor, while a polyphyletic group does not contain the common ancestor at all.
For example, “non-flying mammals” is paraphyletic because it does not include bats, which share a common ancestor with non-flying mammals. “Flying vertebrates” is polyphyletic because it does not include the LCA of vertebrates, which did not fly.
In order to ensure that all resulting groups are monophyletic, the characteristics used in a cladistic analysis are described relative to the group under study by the terms plesiomorphy, apomorphy, and synapomorphy. A plesiomorphy is a characteristic that arose before the group’s last common ancestor (LCA), an apomorphy is one that arose after the LCA, and a synapomorphy is one that arose in the LCA. An apomorphy can be distinguished because it will be present in only some members of the group. Plesiomorphies and synapomorphies are present in all members of the group; they can be distinguished by comparing the group to other groups (outgroups). A plesiomorphy will be found in at least one outgroup, while a synapomorphy will not be present in any outgroup.
Once these determinations have been made, researchers can use computer software to generate a cladogram of the group under study.
Applying Bioinformatics: Sequence Analysis
The first cladistic analyses had to rely on the appearance of organisms, or morphology. Molecular systematics uses the properties of biochemical molecules unique to each species as the characteristics of a cladistic analysis. In other words, DNA, RNA, and protein sequencing are used to ferret out useful characteristics for analysis. This relatively new method can only be used on extant (and possibly recently extinct) species, since no biochemical information is available for fossils.
Nevertheless, in the short time this technique has been available, it has proven itself invaluable in biological classification, especially for organisms with relatively few morphological characteristics, such as bacteria.
Ribosomal RNA changes more slowly than any other part of the genome. Based on a cladistic analysis of rRNA, microbiologist Carl Woese discovered that certain prokaryotic organisms are more closely related to eukaryotes (which include plants, animals, and fungi) than to other prokaryotes.
In 1977, he defined a new level of taxonomy above the kingdom (the highest level under the Linnaean system): the domain. He named his newly discovered clade of prokaryotes Archaea and showed that it shares a common ancestor with a second domain, Eukaryota, and that these constitute a clade separate from the third domain, Bacteria. His work illustrates the power of molecular systematics: almost as soon as it was developed as a discipline, it revealed a fundamental but unsuspected fact about the evolution of life on earth.