Phylogeny is used for categorizing organisms. Cladistics is a much more sophisticated and involves studying genetics.
What is phylogeny?
In 1735, the Swedish naturalist Carl von Linné (better known by his Latinized name, Carolus Linnaeus) published an 11-page monograph entitled Systema Naturae, in which he grouped plants and animals into categories according to shared characteristics. In following editions, he refined the system and developed a standard of nomenclature based on it, giving each species its own generic and specific name. Linnaeus’ system was so well-constructed that it remains the standard way of categorizing organisms almost three centuries later.
The prevailing view in Linnaeus’ day was that all species were created instantaneously and ex nihilo by God. Under that idea, the natural hierarchy observed by Linnaeus seemed a peculiar phenomenon. More than a century later, with the publication of Darwin’s theory that all organisms share a common ancestor, the reason for these relationships became apparent: they are family ties. The current understanding of biologists is that the different varieties of life on earth belong to the same family tree — or more descriptively, a family bush. The tracing of these family relationships is called phylogeny.
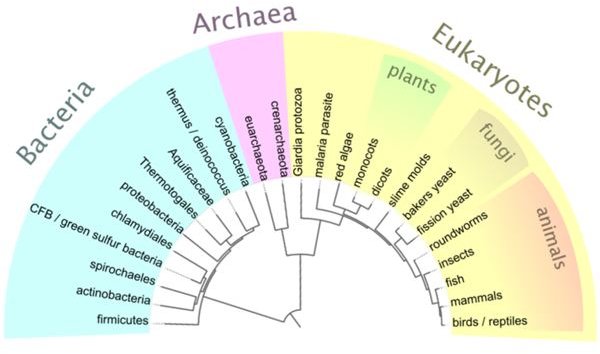
Below: A simplified cladogram illustrating the Tree of Life. Image from Wikimedia Commons by User:McCormack, licensed under the GNU Free Documentation License.
The Tree of Life: A Cladogram
Studying phylogeny: How cladistics works
When grouping related organisms, biologists first identify a number of variable characteristics found in members of the group under study. The process of merely comparing which members share which characteristics is called phenetics. Cladistics is more sophisticated than phenetics because it traces not just similarity, but evolutionary relationships.
A group of organisms sharing a common ancestor is called a clade, from Greek klados, or “branch,” and this term leads to the name cladistics, the study of clades. To trace evolutionary history and distinguish clades, biologists must determine whether each characteristic under study represents an older, more ancestral characteristic (plesiomorphy) or a more recently evolved characteristic (apomorphy). These could be respectively called “primitive” and “advanced,” but because those terms imply value judgments, cladists prefer the terms “basal” and “derived.”
To determine whether characteristics are basal or derived, they are compared to one or more outgroups, that is, groups of organisms outside the group under study. An apomorphy shared by all members of the group but not by outgroups is considered a derived characteristic of the last common ancestor. This type of characteristic is called a synapomorphy.
After identifying apomorphies, synapomorphies, and plesiomorphies, researchers can apply an analysis (typically using a computer) to determine clades. Smaller clades nest within larger clades. In another way of looking at things, smaller clades are always branches from larger clades. The result of a cladistic analysis is a tree diagram called a cladogram.
The role of genetics in cladistics
Before the study of genetics, naturalists had to rely mainly on the appearance of organisms (morphology) to determine phylogenetic relationships. Indeed, when studying extinct species, researchers are still constricted to morphology. For cladistic analysis of extant species whose genomes can be sequenced, however, genetics has powerful advantages over morphology. The use of gene sequences and other biochemical characteristics in cladistics is called molecular systematics.
Because a cladogram maps out evolutionary history, it has an implicit time scale. DNA changes occur at steady rates, a fact which leads to the concept of the “molecular clock.” Using this idea, researchers can determine the relative ages of clades. In some cases, a good approximation of the rate of change of a particular gene is available. In these cases, the clades found in the analysis can be dated in actual years.
Cladistics is the more powerful method yet developed to for tracing ancestral relationships of life on earth. The study of familial relationships using sequence analysis is a major field in the science of bioinformatics.